Building a Variational Autoencoder - Advances in Condition Monitoring, Pt VI

In this post, we’ll explore the variational autoencoder (VAE) and see how we can build one for use on the UC Berkeley milling data set. A variational autoencoder is more expressive than a regular autoencoder, and this feature can be exploited for anomaly detection.
We’ve explored the UC Berkeley milling data set – now it’s time for us to build some models! In part IV of this series, we discussed how an autoencoder can be used for anomaly detection. However, we’ll use a variant of the autoencoder – a variational autoencoder (VAE) – to conduct the anomaly detection.
In this post, we’ll see how the VAE is similar, and different, from a traditional autoencoder. We’ll learn how to implement the VAE and train it. The next post, Part VI, will use the trained VAEs in the anomaly detection process.
The Variational Autoencoder
The variational autoencoder was introduced in 2013 and today is widely used in machine learning applications. The VAE is different from traditional autoencoders in that the VAE is both probabilistic and generative. What does that mean? Well, the VAE creates outputs that are partly random (even after training) and can also generate new data that is like the data it is trained on.
Again, there are excellent explanations of the VAE online – I’ll direct you to Alfredo Canziani’s deep learning course (video below from Youtube).
But here is my attempt at an explanation:
At a high level, the VAE has a similar structure to a traditional autoencoder. However, the encoder learns different codings; namely, the VAE learns mean codings, , and standard deviation codings, . The VAE then samples randomly from a Gaussian distribution, with the same mean and standard deviation created by the encoder, to generate the latent variables, . These latent variables are then “decoded” to reconstruct the input. The figure below demonstrates how a signal is reconstructed using the VAE.
During training, the VAE works to minimize its reconstruction loss (in our case we use binary cross entropy), and at the same time, force a Gaussian structure using a latent loss. The structure is achieved through the Kullback-Leibler (KL) divergence, with detailed derivations for the losses in the original VAE paper.1 The latent loss is as follows:
where is the number of latent variables, and is an adjustable hyper-parameter introduced by Higgens et al.2
Edit: In the original article I had an incorrect equation for the latent loss — I had instead of — and the reconstruction loss was absolute error instead of binary cross entropy. In addition, the notation I used is different from that of Alfredo Canziani — Alfredo’s is very consistent, so watch his videos!
A VAE learns factors, embedded in the codings, that can be used to generate new data. As an example of these factors, a VAE may be trained to recognize shapes in an image. One factor may encode information on how pointy the shape is, while another factor may look at how round it is. However, in a VAE, the factors are often entangled together across the codings (the latent variables). Tuning the hyper-parameter , to a value larger than one, can enable the factors to disentangle such that each coding only represents one factor at a time. Thus, greater interpretability of the model can be obtained. A VAE with a tunable beta is often called a disentangled-variational-autoencoder, or simply, a -VAE.
Data Preparation
Before going any further, we need to prepare the data. Ultimately, we’ll be using the VAE to detect “abnormal” tool conditions, which correspond to when the tool is in a worn. But first we need to label the data.
As shown in the last post, each cut has an associated amount of wear, measured at the end of the cut. We’ll label each cut as either healthy, degraded, or failed according to the amount of wear on the tool. Here’s the schema:
| State | Label | Flank Wear (mm) |
|---|---|---|
| Healthy | 0 | 0 ~ 0.2 |
| Degraded | 1 | 0.2 ~ 0.7 |
| Failed | 2 | > 0.7 |
I’ve created a data prep class that takes the raw matlab files, a labelled CSV (each cut is labelled with the associated flank wear), and spits out the training/validation/and testing data (you can see the entire data_prep.py in the github repository). However, I want to highlight one function in the class that is important; that is, the create_tensor function.
def create_tensor(self, data_sample, signal_names, start, end, window_size, stride=8):
"""Create a tensor from a cut sample. Final tensor will have shape:
[# samples, # sample len, # features/sample]
Parameters
===========
data_sample : ndarray
single data sample (individual cut) containing all the signals
signal_names : tuple
tuple of all the signals that will be added into the tensor
start : int
starting index (e.g. the first 2000 "samples" in the cut may be from
when the tool is not up to speed, so should be ignored)
end : int
ending index
window_size : int
size of the window to be used to make the sub-cuts
stride : int
length to move the window at each iteration
Returns
===========
c : ndarray
array of the sub-cuts
"""
s = signal_names[::-1] # only include the six signals, and reverse order
c = data_sample[s[0]].reshape((9000, 1))
for i in range(len(s)):
try:
a = data_sample[s[i + 1]].reshape((9000, 1)) # reshape to make sure
c = np.hstack((a, c)) # horizontal stack
except:
# reshape into [# samples, # sample len, # features/sample]
c = c[start:end]
c = np.reshape(c, (c.shape[0], -1))
dummy_array = []
# fit the strided windows into the dummy_array until the length
# of the window does not equal the proper length
for i in range(c.shape[0]):
windowed_signal = c[i * stride : i * stride + window_size]
if windowed_signal.shape == (window_size, 6):
dummy_array.append(windowed_signal)
else:
break
c = np.array(dummy_array)
return cThe create_tensor function takes an individual cut, breaks it up into chunks, and puts them into a single array. It breaks the cut signal up into chunks using a window of a fixed size (the window_size variable) and then “slides” the window along the signal. The window “slides” by a preditermined amount, set by the stride variable.
We’ll take each of the 165 cuts (remember, two cuts from the original 167 are corrupted) and apply a window size of 64 and a stride of 64 (no overlap between windows). I’ve visually inspected each cut and selected when the “stable” cutting region occurs, which is usually five seconds or so after the signal begins collection, and a few seconds before the signal collection ends. This information is stored in the “labels_with_tool_class.csv” file.
With all that, we can then create the training/validation/testing data sets. Here is what the script looks like:
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from pathlib import Path
from sklearn.model_selection import train_test_split
import data_prep
data_file = Path("mill.mat")
# helper functions needed in the data processing
def scaler(x, min_val_array, max_val_array):
'''Scale an array across all dimensions'''
# get the shape of the array
s, _, sub_s = np.shape(x)
for i in range(s):
for j in range(sub_s):
x[i,:,j] = np.divide((x[i,:,j] - min_val_array[j]), np.abs(max_val_array[j] - min_val_array[j]))
return x
# min-max function
def get_min_max(x):
'''Get the minimum and maximum values for each
dimension in an array'''
# flatten the input array http://bit.ly/2MQuXZd
flat_vector = np.concatenate(x)
min_vals = np.min(flat_vector,axis=0)
max_vals = np.max(flat_vector,axis=0)
return min_vals, max_vals
# use the DataPrep module to load the data
prep = data_prep.DataPrep(data_file)
# load the labeled CSV (NaNs filled in by hand)
df_labels = pd.read_csv("labels_with_tool_class.csv")
# Save regular data set. The X_train, X_val, X_test will be used for anomaly detection
# discard certain cuts as they are strange
cuts_remove = [17, 94]
df_labels.drop(cuts_remove, inplace=True)
# use the return_xy function to take the milling data, select the stable cutting region,
# and apply the window/stride
X, y = prep.return_xy(df_labels, prep.data,prep.field_names[7:],window_size=64, stride=64)
# use sklearn train_test_split function
# use stratify to ensure that the distribution of classes is equal
# between each of the data sets
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.33, random_state=15, stratify=y)
X_val, X_test, y_val, y_test = train_test_split(X_test, y_test, test_size=0.50, random_state=10, stratify=y_test)
# get the min/max values from each dim in the X_train
min_vals, max_vals = get_min_max(X_train)
# scale the train/val/test with the above min/max values
X_train = scaler(X_train, min_vals, max_vals)
X_val = scaler(X_val, min_vals, max_vals)
X_test = scaler(X_test, min_vals, max_vals)
# training/validation of VAE is only done on healthy (class 0)
# data samples, so we must remove classes 1, 2 from the train/val
# data sets
X_train_slim, y_train_slim = prep.remove_classes(
[1,2], y_train, X_train
)
X_val_slim, y_val_slim = prep.remove_classes(
[1,2], y_val, X_val
)The final distribution of the data is shown below. As noted in Part IV of this series, we’ll be training the VAE on only the healthy data (class 0). However, checking the perfomance anomaly detection will be done using all the data. In other words, we’ll be training our VAE on the “slim” data sets.
| Data Split | No. Sub-Cuts | Healthy % | Degraded % | Failed % |
|---|---|---|---|---|
| Training full | 11,570 | 36.5 | 56.2 | 7.3 |
| Training slim | 2,831 | 100 | - | - |
| Validation full | 1,909 | 36.5 | 56.2 | 7.3 |
| Validation slim | 697 | 100 | - | - |
| Testing full | 1,910 | 36.5 | 56.2 | 7.3 |
Building the Model
We now understand what a variational autoencoder is and how the data is prepared. Time to build!
Our VAE will be made up of layers consisting of convolutions layers, batch normalization layers, and max pooling layers. The figure below shows what one of our VAE models could look like.
I won’t be going through all the details of the model — you can look at the Jupyter notebook and read my code comments. However, here are some important points:
- I’ve used the temporal convolutional network as the basis for the convolutional layers. The implementation is from Philippe Remy — thanks Philippe! You can find his github repo here.
- Aurélien Geron’s book, “Hands-On Machine Learning with Scikit-Learn, Keras, and TensorFlow”, is great. In particular, his section on VAEs was incredibly helpful, and I’ve used some of his methods here. There is a Jupyter notebook from that section of his book on his github. Thanks Aurélien! 3
- I’ve used rounded accuracy to measure how the model performs during training, as suggested by Geron.
Here is, roughly, what the model function looks like. Note, I’ve also included the Sampling class and rounded accuracy function.
import tensorflow as tf
from tensorflow import keras
from tcn import TCN
import numpy as np
import datetime
class Sampling(keras.layers.Layer):
'''Used to sample from a normal distribution when generating samples
Code from Geron, Apache License 2.0
https://github.com/ageron/handson-ml2/blob/master/17_autoencoders_and_gans.ipynb
'''
def call(self, inputs):
mean, log_var = inputs
return K.random_normal(tf.shape(log_var)) * K.exp(log_var / 2) + mean
def rounded_accuracy(y_true, y_pred):
'''Rounded accuracy metric used for comparing VAE output to true value'''
return keras.metrics.binary_accuracy(tf.round(y_true), tf.round(y_pred))
def model_fit(
X_train_slim,
X_val_slim,
beta_value=1.25,
codings_size=10,
dilations=[1, 2, 4],
conv_layers=1,
seed=31,
start_filter_no=32,
kernel_size_1=2,
epochs=10,
earlystop_patience=8,
verbose=0,
compile_model_only=False,
):
_, window_size, feat = X_train_slim.shape
# save the time model training began
# this way we can identify trained model at the end
date_time = datetime.datetime.now().strftime("%Y%m%d-%H%M%S")
# set random seeds so we can somewhat reproduce results
tf.random.set_seed(seed)
np.random.seed(seed)
end_filter_no = start_filter_no
inputs = keras.layers.Input(shape=[window_size, feat])
z = inputs
#### ENCODER ####
for i in range(0, conv_layers):
z = TCN(
nb_filters=start_filter_no,
kernel_size=kernel_size_1,
nb_stacks=1,
dilations=dilations,
padding="causal",
use_skip_connections=True,
dropout_rate=0.0,
return_sequences=True,
activation="selu",
kernel_initializer="he_normal",
use_batch_norm=False,
use_layer_norm=False,
)(z)
z = keras.layers.BatchNormalization()(z)
z = keras.layers.MaxPool1D(pool_size=2)(z)
z = keras.layers.Flatten()(z)
codings_mean = keras.layers.Dense(codings_size)(z)
codings_log_var = keras.layers.Dense(codings_size)(z)
codings = Sampling()([codings_mean, codings_log_var])
variational_encoder = keras.models.Model(
inputs=[inputs], outputs=[codings_mean, codings_log_var, codings]
)
#### DECODER ####
decoder_inputs = keras.layers.Input(shape=[codings_size])
x = keras.layers.Dense(start_filter_no * int((window_size / (2 ** conv_layers))), activation="selu")(decoder_inputs)
x = keras.layers.Reshape(target_shape=((int(window_size / (2 ** conv_layers))), end_filter_no))(x)
for i in range(0, conv_layers):
x = keras.layers.UpSampling1D(size=2)(x)
x = keras.layers.BatchNormalization()(x)
x = TCN(
nb_filters=start_filter_no,
kernel_size=kernel_size_1,
nb_stacks=1,
dilations=dilations,
padding="causal",
use_skip_connections=True,
dropout_rate=0.0,
return_sequences=True,
activation="selu",
kernel_initializer="he_normal",
use_batch_norm=False,
use_layer_norm=False,
)(x)
outputs = keras.layers.Conv1D(
feat, kernel_size=kernel_size_1, padding="same", activation="sigmoid"
)(x)
variational_decoder = keras.models.Model(inputs=[decoder_inputs], outputs=[outputs])
_, _, codings = variational_encoder(inputs)
reconstructions = variational_decoder(codings)
variational_ae_beta = keras.models.Model(inputs=[inputs], outputs=[reconstructions])
latent_loss = (
-0.5
* beta_value
* K.sum(
1 + codings_log_var - K.exp(codings_log_var) - K.square(codings_mean),
axis=-1,
)
)
variational_ae_beta.add_loss(K.mean(latent_loss) / (window_size * feat))
variational_ae_beta.compile(
loss="binary_crossentropy", optimizer="adam", metrics=[rounded_accuracy]
)
# count the number of parameters so we can look at this later
# when evaluating models
param_size = "{:0.2e}".format(
variational_encoder.count_params() + variational_decoder.count_params()
)
# Save the model name
# b : beta value used in model
# c : number of codings -- latent variables
# l : numer of convolutional layers in encoder (also decoder)
# f1 : the starting number of filters in the first convolution
# k1 : kernel size for the first convolution
# k2 : kernel size for the second convolution
# d : whether dropout is used when sampling the latent space (either True or False)
# p : number of parameters in the model (encoder + decoder params)
# eps : number of epochs
# pat : patience stopping number
model_name = (
"TBVAE-{}:_b={:.2f}_c={}_l={}_f1={}_k1={}_dil={}"
"_p={}_eps={}_pat={}".format(
date_time,
beta_value,
codings_size,
conv_layers,
start_filter_no,
kernel_size_1,
dilations,
param_size,
epochs,
earlystop_patience,
)
)
# use early stopping
# stop training model when validation error begins to increase
earlystop_callback = tf.keras.callbacks.EarlyStopping(
monitor="val_loss",
min_delta=0.0002,
patience=earlystop_patience,
restore_best_weights=True,
verbose=1,
)
# fit model
history = variational_ae_beta.fit(
X_train_slim,
X_train_slim,
epochs=epochs,
batch_size=256,
shuffle=True,
validation_data=(X_val_slim, X_val_slim),
callbacks=[earlystop_callback],
verbose=verbose,
)
return date_time, model_name, history, variational_ae_beta, variational_encoderTraining the Model
Time to begin training some models. To select the hyperparameters, we’ll be using a random search. Why a random search? Well, it’s fairly simple to implement and has been shown to yield better results when compared to a grid search.4 Scikit-learn has some nice methods for implementing a random search, and that’s what we’ll use.
We’ll be training a bunch of different VAEs, all with different parameters. After each VAE has been trained (trained to minimize reconstruction loss) and the model saved, we’ll go through the VAE model and see how it performs in anomaly detection. Here’s a diagram of the random search training process:
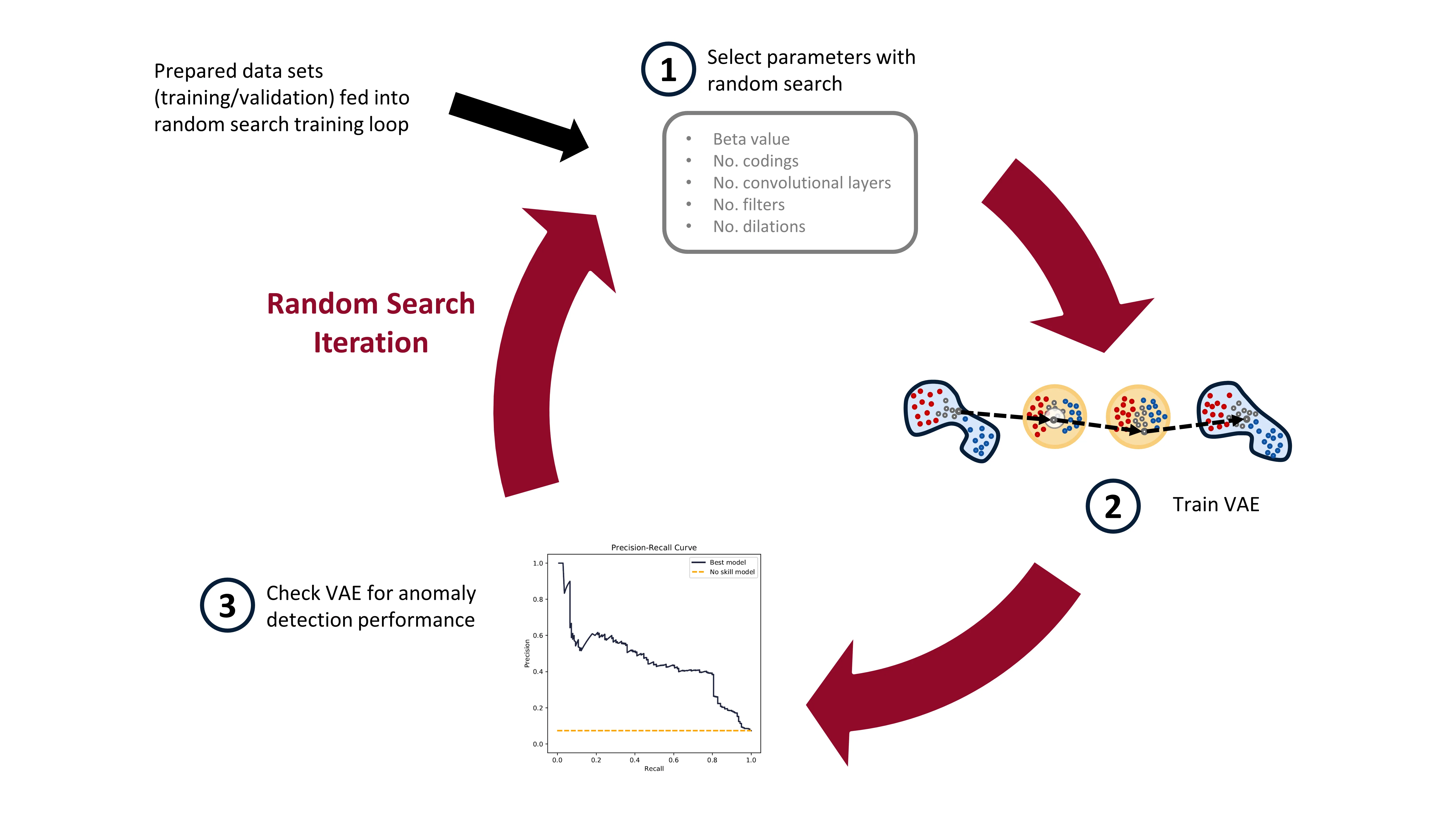
In practice, when I ran this experiment, I trained about 1000 VAE models on Google Colab (yay free GPUs!). After all 1000 models were trained, I moved them to my local computer, with a less powerful GPU, and then checked the models for anomaly detection performance. Google Colab has access to great GPUs, but your time with them is limited, so it makes sense to maximize the use of the GPUs on them this way. I’ll detail how the trained models are checked for anomaly detection performance in the next blog post.
I’m not going to show the training loop code here. You can see it all in the Google Colab notebook, or on the github.
Conclusion
In this post we’ve learned about the VAE; prepared the milling data; and implemented the training of the VAE. Phew! Lots of work! In the next post, we’ll evaluate the trained VAEs for anomaly detection performance. I’ll also explain how we use the precision-recall curve, and we’ll make some pretty graphics (my favourite!). Stay tuned!
Footnotes
-
Kingma, Diederik P., and Max Welling. “Auto-encoding variational bayes.” arXiv preprint arXiv:1312.6114 (2013). ↩
-
Higgins, Irina, et al. “beta-vae: Learning basic visual concepts with a constrained variational framework.” (2016). ↩
-
Géron, Aurélien. Hands-on machine learning with Scikit-Learn, Keras, and TensorFlow: Concepts, tools, and techniques to build intelligent systems. O’Reilly Media, 2019. ↩
-
Bergstra, James, and Yoshua Bengio. “Random search for hyper-parameter optimization.” The Journal of Machine Learning Research 13.1 (2012): 281-305. ↩